

While the majority of proteins spend most of their time folded in a stable three-dimensional structure, a particular class of proteins known as intrinsically disordered proteins lacks a stable fold. Several of these proteins have been found to be associated with disease, and they may be useful as well as models of the unfolded protein state, for investigating the first steps in the protein folding process. Parkinson’s disease is a neurodegenerative disorder that leads to motor and cognitive impairment, generally diagnosed after the age of 60, for which there is currently no cure. The intrinsically-disordered protein α-synuclein is the primary component of protein aggregates known as Lewy bodies found in the neurons of Parkinson’s patients, and genetic mutations causing either a change in α-synuclein’s sequence or excessive production of the protein have been linked to disease, suggesting that α-synuclein plays a key role in disease development and that finding drugs to interfere with its abnormal activity may be a promising strategy for stopping or slowing disease progression.
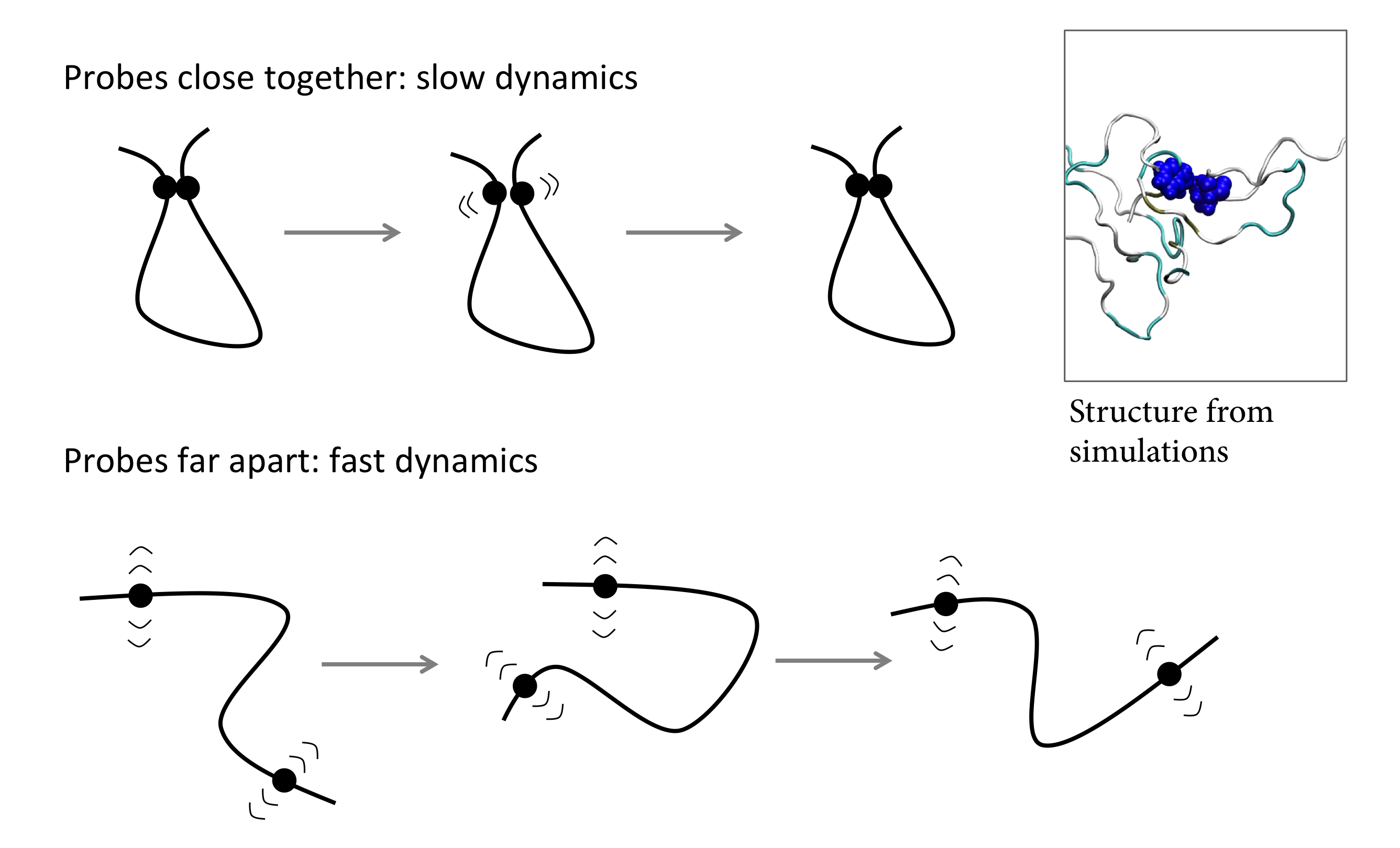
In collaboration with Lisa Lapidus’ group at MSU, we aim to study α-synuclein at a detailed physical level using a combination of theory, simulation, and experiment. Using established theory and two experimental techniques: Förster resonance energy transfer and tryptophan-cysteine quenching, our results suggest that two amino acids of a protein diffuse more slowly relative to one another when they are close together in space. Molecular dynamics simulations of WT α-synuclein using a recently developed force field reproduce this result and provide a possible explanation: clusters of amino acids form transiently, causing amino acids within the protein to sometimes get stuck together. We think these clusters could be important for nucleating protein aggregation and therefore may be relevant to disease. We believe our results may be applicable generally to intrinsically-disordered proteins and the disordered protein state.
A 2-μs trajectory of WT alpha synuclein is available under the list of publications on our website.

| 2018 | ||
| 2. | Jaie Woodard, Kinshuk R. Srivastava, Gil Rahamim, Asaf Grupi, Steven Hogan, David J. Witalka, Grzegorz Nawrocki, Elisha Haas, Michael Feig, Lisa J. Lapidus: Intramolecular Diffusion in α-Synuclein: It Depends on How you Measure It Biophysical Journal (2018) 115, 1190-1199 Abstract PDF | |
| 2017 | ||
| 1. | Jing Huang, Sarah Rauscher, Grzegorz Nawrocki, Ting Ran, Michael Feig, Bert L. de Groot, Helmut Grubmüller, Alexander D. MacKerell Jr.: CHARMM36m: An Improved Force Field for Folded and Intrinsically Disordered Proteins Nature Methods (2017) 14, 71-73 Abstract PDF |